 
The markersize is only the width (not the area) of the marker: ax. from adjustText import adjusttext define criteria for filtering the data minscaledrank 0 maxscaledrank 5 minbackwin 1. If even gets more confusing if you use plot (which is faster if all markers should have the same size as the docs state as "Notes"). #ax.plot(x_data, y_data, "o",markersize=2*r_)Īs soon as you add an edge (so a non-zero border around the markers), they will overlap: So calculating the area is not pi * r_**2 but rather a square: (2*r_)**2 # open figureĪx.scatter(x_data, y_data, s=marker_size,linewidths=1) one would never calculate the actual area of the symbol). We are not talking about the area of the symbol (in this case a circle) but of a bounding box of the marker (imagine, you want to control the size of a star or an asterix as marker. This is due a misunderstanding of the area of the markers. This is a rather tedious idea, because you need to plot the data twice but you keep the autosizing of the axes. Scl = ax.get_xlim() - ax.get_xlim()Īx.scatter(x_data, y_data, s=marker_size/scl**2, edgecolors='blue',color='red') or scale the markers's size according to the new limits (you will need to know them or do the plotting again) # plot with invisible colorĪx.scatter(x_data, y_data, s=marker_size, color=(0,0,0,0)) R_ = ax.ansform() - ax.ansform() # points # create a n x n square with a marker at each point as dummy data The complete code is: import matplotlib.pyplot as plt So either set the limits to the known size before plotting: ax.set_xlim((0,n-1)) but you are not reducing the marker-size. The bottom line is that you determine the coordinate transformation with the default axis-limits ( (0,1) x (0,1)) and enlarges them afterwards to (-0.75, 15.75)x (-0.75, 15.75). Yes, the problem is that you determine the coordinate-units-to-figure-points size before plotting, so before setting the limits, which influence the coordinate-units but not the overall figure size. The radius is (almost) exactly the r=0.5 coordinate-units that you wanted to have. You can see this if you plot just 4 points ( n=2): When I run it with s=r_ I get the result on the left and with s=marker_size I get the result on the right of the following image: R_ = ax.ansform() - ax.ansform()Īx.scatter(x_data, y_data, s=marker_size, edgecolors='black') 
# calculate the marker size so that the markers touch # create a n x n square with a marker at each point Here is an example code of what I want to accomplish, with the results as an image below the code: import matplotlib.pyplot as plt It could also be that running matplotlib from WSL via VcXsrv is causing this problem. I tried transforming the given radius into points via ansData and then calculating the area via pi * r^2, but I did not succeed. I've read in pyplot scatter plot marker size, that the marker size is given as the area of the marker in points^2. Plt.I want to have the markers of a scatter plot match a radius given in the data coordinates. # compare with no borders, and denser hatch. 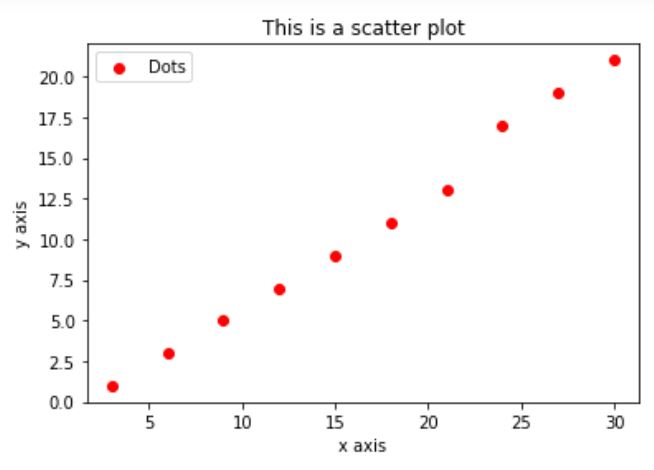
Plt.scatter(x,y, s=500, marker='s', edgecolor='black', linewidth=3, facecolor='green', hatch='|') See the code example below to produce scatter plots such as these: However, I don't think the thickness of individual lines within hatching is controllable. You can increase the density of hatching, by repeating symbols (in the example below, the '|' is repeated in the R/H pane note that to obtain NW->SE diagonal lines the symbol must be escaped so needs twice as many characters to really double it - '\' is density 2 while '||||' is density 4).To control the marker border thickness, you just need to set the linewidth for scatter() (or markeredgewidth for plot()). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
